Project background
The McMullan team study the co-evolution of plants and their pathogens, with a particular focus on wild crop relatives (non-domesticated plant species related to crops).
Our team use phylogenomics, population genomics, and transcriptomics to understand pathogen adaptation, gene flow, and population structure across natural landscapes. Agriculture, which arose only ~10,000 years ago, provides a powerful natural perturbation of pre-existing host-pathogen systems; a natural experiment allowing us to study pathogen adaptation and speciation onto domesticated hosts, and the spillback consequences for the wild plant communities these pathogens continue to inhabit. We focus on two systems which were domesticated at very different times, wheat (early) and sugar beet (late), and we offer two projects, giving you the flexibility to tailor your training.
Project 1: Understanding Gaeumannomyces tritici ecology across natural grass hosts
We aim to understand how Gaeumannomyces tritici moves between hosts in the soil. This fungus is the most devastating root disease of wheat worldwide, and it persists in soil on debris between crop seasons. Rotation is currently a key defence strategy. However, this fungus existed long before wheat cultivation, and we aim to better understand the ecological conditions under which it evolved 10,000 years ago, before wheat was a crop. Understanding this adaptation and the evolution of host range is central to understanding both its life history and the evolution of virulence.
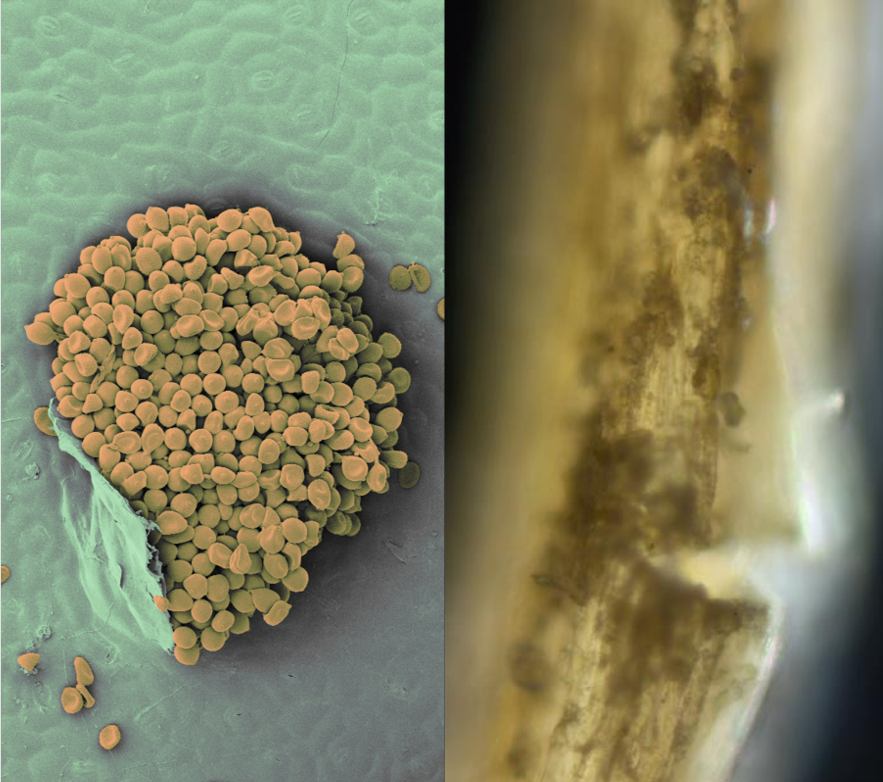
Using established infection protocols, you will culture G. tritici on wheat and related wild grasses, then apply complementary approaches to quantify fungal presence: qPCR for total fungal biomass, microscopy for structural analysis, and newly developed single nuclei extraction methodologies for propagule counting. Results will shed light on fungal lifecycle, and improve our understanding of pathogenicity and resistance.
See Hill, R. et al. Evolutionary genomics reveals variation in structure and genetic content implicated in virulence and lifestyle in the genus Gaeumannomyces. BMC Genomics 26, 239 (2025).
Project 2: Wild resistance gene – effector binding complementarity
Do resistance genes in wild hosts recognise rust effectors from crop-adapted pathogenic strains? Wild crop relatives can serve as a reservoir for pathogens that can go on to infect crops, and vice versa. Sugar beet (Beta vulgaris) is one of the most recently domesticated crops, bred in Europe and still grown alongside its wild crop relative: sea beet. Sugar beet and sea beet share many pathogen species, but whereas sugar beet varieties have been bred for agronomic traits, sea beet still contains substantial genetic diversity. Are those wild resistance genes shaped by co-evolutionary dynamics with locally adapted pathogen populations?
We have found beet rust living on both wild and crop beets, and that alleles appear to be differentially favoured in the pathogens living on wild and crop populations. We have also identified resistance genes in wild beets that appear to be favoured in regions near to where sugar beet is grown. We are now interested in using the predicted 3D structure of these resistance genes and effectors, to identify evidence for their interaction. Results will shed light on whether wild hosts and crop pathogen populations are already locked in a molecular arms race, a question with broad implications for understanding coevolution across domesticated landscapes.
See McMullan, M. et al. Developing a crop- wild-reservoir pathogen system to understand pathogen evolution and emergence. eLife 14, e91245 (2025).